Why high Tm LNA® probes improve diagnostic accuracy?
In molecular diagnostics, probe design directly impacts sensitivity and specificity. One of the key innovations in this space is the use of LNA® probes, known for their high melting temperature (Tm) and ability to function effectively even at short lengths. But why does a higher Tm matter?
What is a LNA® probe?
LNA® probes are oligonucleotides that incorporate Locked Nucleic Acid (LNA®) bases, a class of chemically modified nucleotides. In LNA®, the ribose structure is “locked” by a bridge between the 2’-O and 4’-C atoms (Obika et al. 1997, Suresh et al. 2013). This modification constrains the nucleotide into a rigid conformation that enhances hybridization properties (see Figure 1).
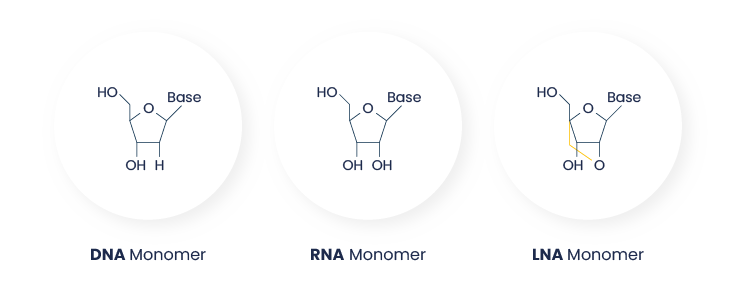
Figure 1: Structural difference between DNA, RNA and LNA® nucleotides. The 2’-O–4’-C methylene bridge in LNA® locks the ribose in a C3’-endo conformation (Pande and Nilsson et al. 2008), promoting optimal base stacking and duplex stability, and resulting in increased hybridization affinity and higher Tm.
What is melting temperature (Tm)?
The melting temperature (Tm) is the temperature at which half of a nucleic acid probe is bound to its target sequence and half is unbound. A higher Tm indicates stronger hybridization between the probe and its target sequence.
Tm depends on several factors:
- Sequence composition (GC vs AT content)
- Probe length
- Salt conditions
- Presence of chemical modifications such as LNA®
Why LNA® probes have a higher Tm?
The increased Tm of LNA® probes results from a cascade of structural effects (Figure 2). In LNA®, the ribose ring is chemically constrained by a bridge linking the 2’-O and 4’-C atoms, effectively “locking” the sugar into a fixed conformation. This structural constraint induces conformational rigidity and forces the nucleotide into a C3’-endo geometry (Pande and Nilsson et al. 2008), similar to that found in RNA and optimal for duplex formation.
As a result, the bases are more precisely aligned, leading to improved base stacking and stronger π–π interactions between adjacent nucleotides (Petersen et al. 2000, Vester and Wengel et al. 2004). These structural improvements enhance the overall stability of the probe–target duplex, resulting in stronger hybridization and increased binding affinity.
This increased stability directly translates into a higher melting temperature (Tm) (Lundin et al. 2013), meaning that more energy is required to separate the probe from its target sequence. In practice, the incorporation of a single LNA® base can increase the Tm of the probe–target duplex by approximately 2–8°C, depending on sequence context (Suresh et al. 2013, Pande and Nilsson et al. 2008).
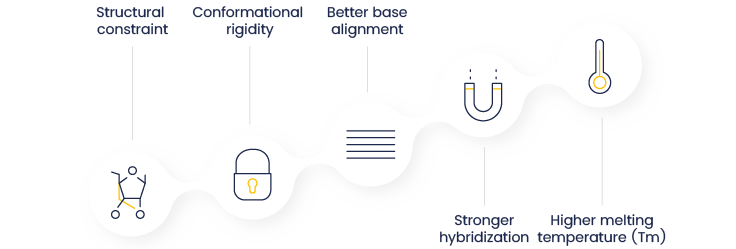
Figure 2. How comes LNA®-modified probes have a higher Tm. LNA® bases introduce a chemical bridge that locks the ribose structure, enforcing a rigid conformation and improving base stacking. This increases duplex stability and binding affinity, leading to a higher melting temperature (Tm).
Why a higher Tm is an advantage in diagnostics?
1. Increased hybridization stability
High-Tm probes remain strongly bound to their target sequences, even under stringent assay conditions such as those used in PCR or hybridization assays. This enhanced stability ensures more efficient target capture and reduces the likelihood of probe dissociation during analysis (Vester and Wengel et al. 2004). As a result, assays benefit from improved sensitivity and more reliable detection, particularly when working with AT-rich or low-abundance targets.
2. Superior specificity and mismatch discrimination
One of the most powerful advantages of LNA® probes is their ability to increase the contrast between perfectly matched and mismatched sequences (You et al. 2006, Owczarzy et al. 2011). With standard DNA probes, mismatches are often only slightly destabilizing, making discrimination difficult. In contrast, LNA® probes strongly stabilize perfectly matched duplexes while significantly destabilizing mismatched ones. This effect arises because LNA® enforces an optimal geometry for correct base pairing, while mismatches become structurally unfavorable within the rigid conformation. Consequently, the difference in melting temperature (ΔTm) between matched and mismatched targets is increased. At elevated temperatures, perfectly matched sequences remain bound, whereas mismatches dissociate, enabling highly accurate detection of SNPs (Mouritzen et al. 2003, Johnson et al. 2004), point mutations (Pu and Wu et al. 2025), and genetic variants.
3. Reduced background noise
The high melting temperature of LNA® probes allows assays to be performed under more strin-gent experimental conditions, where weak or non-specific interactions are minimized or eliminated (Pena et al. 2009). By reducing non-specific binding, these conditions improve the clarity of the signal and enhance the overall signal-to-noise ratio. This leads to cleaner assay outputs and more confident interpretation of results.
4. Enables shorter, more specific probes
Unlike standard DNA probes, which lose stability when shortened, LNA®-modified probes retain a high melting temperature even at reduced lengths (Vester and Wengel et al. 2004). This property enables the design of shorter probes that maintain strong hybridization while improving target accessibility (Ugozzoli et al. 2004). As a result, shorter LNA® probes can more effectively target small or structured sequences, reduce off-target interactions, and increase overall assay precision. Therefore, LNA® is recommended for use in any hybridization assay that requires high specificity like custom oligonucleotides for PCR-based applications , double dye probes , FISH probes , or molecular beacons .
5. Robust performance in complex samples
Clinical and biological samples, such as blood or FFPE tissue biopsies (Pena et al. 2009), often contain degraded nucleic acids, inhibitors, and complex matrices that can interfere with hybridization. High-Tm LNA® probes maintain stable binding under these challenging conditions, contributing to consistent assay performance. In addition, LNA®-modified oligonucleotides exhibit increased resistance to nuclease degradation compared to standard DNA probes, further supporting their reliability in living cells environments (Riahi et al. 2013).
Techniques where LNA® probes are particularly valuable
Thanks to their high affinity, elevated melting temperature (Tm), and strong mismatch discrimination capabilities, LNA®-modified probes are compatible with a wide range of nucleic acid-based technologies. Their ability to maintain stable and specific hybridization, even with short or challenging targets, makes them particularly valuable in applications requiring high sensitivity and precision.
LNA® probes are commonly used in:
- PCR, RT-PCR, and real-time PCR (qPCR) for sensitive and specific target detection
- Digital PCR (dPCR) for rare mutation and low-copy target analysis
- Microarrays and capture probe technologies to improve hybridization performance and target discrimination
- Fluorescent in situ hybridization (FISH/ISH) for tissue-based detection and spatial localization of nucleic acids
- microRNA profiling and detection, where short target sequences require enhanced duplex stability
- Emerging CRISPR-based diagnostic systems, where improved specificity is increasingly important
Beyond diagnostics, LNA® technology has also been widely explored in antisense oligonucleotides and gene silencing approaches because of its high affinity, nuclease resistance, and improved target specificity (Wahlestedt et al. 2000, Bengtson Løvendorf et al. 2023).
Key diagnostic applications of LNA® primers and probes
Table 1. Key diagnostic applications of high-Tm LNA® primers and probes.
| Diagnostic applications | Examples | Scientific references |
| microRNA detection (very short targets) | miR-21, cancer prognostic biomarker (ISH) | James et al. 2020 |
| miR panel (Microarray platform for genome-wide profiling of mature miRNAs [miChip]) | Castoldi et al. 2006 | |
| Plant miRNA (Northern blot analysis) | Valoczi et al. 2004 | |
| SNP detection, mutation analysis, variant discrimination | Allele-specific PCR assays within the cystic fibrosis (CFTR) gene | Latorra et al. 2003 |
| Multiplex detection | 14-plex respiratory virus panel (Nested real-time PCR) | Zhao et al. 2020 |
| Rice GMO analysis (ULNA-ddPCR) | Yang et al. 2021 | |
| CRISPR-based detection (emerging) | CRISPR diagnostics (e.g., CALSA) on ssDNA or genomic DNA | Sun et al. 2024 |
| Viral variant detection, pathogen genotyping | Liu et al. 2025 | |
| Circulating free DNA (cfDNA), rare target detection | PNA-LNA clamp PCR on liquid biopsy (ctDNA) to detect lung cancer biomarkers (e.g., EGFR) | Vendrell et al. 2017 |
| Diagnostics in biological tissues | miRNA detection in FFPE tissue samples (ISH) | Pena et al. 2009 |
| Diagnostics in living cells | Intracellular mRNA quantification in living cells (single-cell gene expression analysis) | Riahi et al. 2013 |
Conclusions
A high melting temperature (Tm) is not just a technical parameter, it is a key driver of diagnostic performance.
By enabling stronger and more stable binding, greater specificity, and the use of shorter probes, LNA® probes provide a powerful solution for detecting small, rare, or discriminating highly similar nucleic acid sequences with high precision.
However, their increased affinity and elevated Tm also require careful design and optimization. As a result, LNA® probes are best suited for high-value diagnostic assays, where improved performance justifies the added complexity and cost.
At Eurogentec, we support these advanced applications by offering custom LNA®-modified probes, tailored to your specific assay requirements. To streamline your workflow, these probes can be easily designed and ordered through our online configurator .
Need help in LNA® probe design?
Diagnostic or testing professionals?
Discover our custom oligo for diagnostic and testing
How to improve bacterial detection by qPCR ?
Discover Takyon™ Etheris a reliable master mix for consistent, sensitive and specific results in microbiology.
Streamline your diagnostic workflow
Outsource your in-house assay preparation and increase manufacturing consistency
References
Bengtson Løvendorf, M., Holm, A. , Petri A. , Albæk Thrue, C. , Uchida, S. , T Venø, M., Kauppinen, S. Knockdown of Circular RNAs Using LNA-Modified Antisense Oligonucleotides. Nucleic Acid Ther. 2023 Jan;33(1):45-57 (2023).
Castoldi, M., Schmidt, S., Benes, V., Noerholm, M., Kulozik, A. E., Hentze, M. W., & Muckenthaler, M. U. A sensitive array for microRNA expression profiling (miChip) based on locked nucleic acids (LNA). RNA, 12(5), 913–920 (2006).
James, V., et al. MicroRNA in situ hybridization in paraffin-embedded cultured cells and tissues using LNA probes. Methods in Molecular Biology, 2148, 73–87 (2020).
Johnson, M. P., Haupt, L. M., & Griffiths, L. R. Locked nucleic acid (LNA) single nucleotide polymorphism (SNP) genotype analysis and validation using real-time PCR. Nucleic Acids Research, 32(6), e55 (2004).
Latorra, D. et al., Enhanced allele-specific PCR discrimination in SNP genotyping using 3' locked nucleic acid (LNA) primers. Hum Mutat. Jul;22(1):79-85 (2003).
Liu, X., et al. Advanced nucleic acid probes for enhanced CRISPR diagnostics. TrAC Trends in Analytical Chemistry, 184, 118496 (2025).
Lundin, K. E., et al. Biological activity and biotechnological aspects of locked nucleic acids. Advanced Genetics, 82, 47–107 (2013).
Mouritzen, P., Nielsen, A. T., Pfundheller, H. M., Choleva, Y., Kongsbak, L., & Møller, S. Single nucleotide polymorphism genotyping using locked nucleic acid (LNA). Expert Review of Molecular Diagnostics, 3(1), 27–38 (2003).
Obika, S., Nanbu, D., Hari, Y., Morio, K., In, Y., Ishida, T., & Imanishi, T. Synthesis of 2′-O,4′-C-methyleneuridine and -cytidine. Novel bicyclic nucleosides having a fixed C3′-endo sugar puckering. Tetrahedron Letters, 38(50), 8735–8738 (1997).
Owczarzy, R., et al. Predicting stability of DNA duplexes in solutions containing magnesium and monovalent cations. Biochemistry, 50(43), 9352–9367 (2011).
Pande, V., & Nilsson, L. Insights into the thermodynamics of LNA hybridization from molecular dynamics simulations. Nucleic Acids Research, 36(5), 1508–1516 (2008).
Pena, J. T. G., et al. miRNA in situ hybridization in formaldehyde and EDC-fixed tissues. Nature Methods, 6(2), 139–141 (2009).
Petersen, M., Bondensgaard, K., Wengel, J., & Jacobsen, J. P. Locked nucleic acid (LNA) recognition of RNA: NMR solution structures of LNA:RNA hybrids. Journal of the American Chemical Society, 122(40), 9932–9941 (2000).
Pu, X., & Wu, S. Recent advances in CRISPR/Cas-based biosensing using modified nucleic acid probes. Biosensors, 15(2), 112 (2025).
Riahi, R., Dean, Z., Wu, T. H., Teitell, M. A., Chiou, P. Y., Zhang, D. D., & Wong, P. K. Detection of mRNA in living cells by double-stranded locked nucleic acid probes. Analyst, 138(17), 4777–4785 (2013).
Sun, Y., et al. An autocatalytic CRISPR-Cas amplification effect propelled by the LNA-modified split activators for DNA sensing. Nucleic Acids Research, 52(7), e39 (2024).
Suresh, M., et al. Locked nucleic acid applications in diagnostics and therapeutics. Journal of Nucleic Acids, 2013, 1–14 (2013).
Ugozzoli, L., et al.LNA and MGB probes in SNP genotyping and mutation analysis. BioTechniques, 36(1), 62–67 (2004).
Valoczi, A., Hornyik, C., Varga, N., Burgyán, J., Kauppinen, S., & Havelda, Z. Sensitive and specific detection of microRNAs by northern blot analysis using LNA-modified oligonucleotide probes. Nucleic Acids Research, 32(22), e175 (2004).
Vendrell, J. A., et al. Detection of EGFR mutations in circulating tumor DNA by PNA-LNA clamp PCR. Scientific Reports, 7, 41445 (2017).
Vester, B., & Wengel, J. LNA (locked nucleic acid): high-affinity targeting of complementary RNA and DNA. Biochemistry, 43(42), 13233–13241 (2004).
Wahlestedt, C., et al. Potent and nontoxic antisense oligonucleotides containing locked nucleic acids. Proceedings of the National Academy of Sciences USA, 97(10), 5633–5638 (2000).
Yang, L., et al. Detection of genetically modified rice using ultra-sensitive LNA-ddPCR. Food Chemistry, 343, 128470(2021).
You, Y., et al. LNA probe-based assay improves mismatch discrimination for SNP genotyping. Nucleic Acids Research, 34(8), e60 (2006).
Zhao, X., et al.Development and evaluation of a multiplex one-tube nested real-time PCR assay using LNA-modified primers for respiratory viruses. Journal of Medical Virology, 92(10), 2120–2128 (2020).